DBTMEE:0016228
| Gene | Pias2 |
| Synonyms | ARIP3
Dib
Miz1
PIASxalpha
PIASxb
PIASxbeta |
| Description | protein inhibitor of activated STAT 2 |
| Locus | NM_001164169 UCSC: chr18:77303947-77385678(+) OLP: chr18:77303947-77385678(+)
NM_001164167 UCSC: chr18:77303947-77394447(+) OLP: chr18:77303947-77394447(+)
NM_008602 UCSC: chr18:77303947-77394447(+) OLP: chr18:77303947-77394447(+)
NM_001164170 UCSC: chr18:77304419-77385678(+) OLP: chr18:77304419-77385678(+)
NM_001164168 UCSC: chr18:77304419-77394447(+) OLP: chr18:77304419-77394447(+)
|
| DB links |
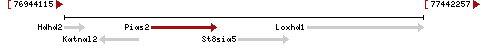
Entrez 17344
MGI 1096566
Ensembl ENSMUSG00000025423
|
| Feature | protein-coding |
| GO_BP |
-
 12 Terms
transcription, DNA-dependent (GO:0006351)
regulation of transcription, DNA-dependent (GO:0006355)
regulation of transcription from RNA polymerase II promoter (GO:0006357)
metabolic process (GO:0008152)
protein sumoylation (GO:0016925)
androgen receptor signaling pathway (GO:0030521)
regulation of osteoblast differentiation (GO:0045667)
positive regulation of transcription, DNA-dependent (GO:0045893)
positive regulation of transcription from RNA polymerase II promoter (GO:0045944)
positive regulation of dendrite morphogenesis (GO:0050775)
regulation of androgen receptor signaling pathway (GO:0060765)
negative regulation of androgen receptor signaling pathway (GO:0060766)
|
| GO_MF |
-
 12 Terms
|
| GO_CC |
-
 2 Terms
|
| KEGG |
-
 5 Terms
|
| Ver2_FPKM |
see DBTMEE:2016706 (v2.0)
|
| SPR_Histone_Erkek_5KaroundTSS |
N/A |
| SPR_Histone_Erkek_2KaroundTSS |
N/A |
| SPR_Methy_Kobayashi_5KaroundTSS |
N/A |
| SPR_Methy_Kobayashi_2KaroundTSS |
N/A |
| SOLiD_OocyteTo4C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | SPR_FPKM | Oo_FPKM | 1C_FPKM | 2C_FPKM | 4C_FPKM | MEF_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 29.4054 | 23.3425 | 24.5543 | 16.3279 | 12.8895 | |
|
| SOLiD_Illumina_Oo_1C_Sperm_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | SPR_FPKM | 1C_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 30.746 | 9.50072 | 25.4671 |
|
| SOLiD_Partheno1C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | 1C_FPKM | p1C_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 30.0112 | 23.1081 | 21.8649 |
|
| SOLiD_Illumina_Oo_p1C_Sperm_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | SPR_FPKM | p1C_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 30.7639 | 9.3706 | 23.9807 |
|
| SOLiD_Partheno4C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | 2C_FPKM | 4C_FPKM | p4C_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 22.7334 | 15.7652 | 16.2577 |
|
| Illumina_Oo_2C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | 2C_FPKM |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 72.5043 | 12.1588 |
|
| SOLiD_SC_Oo_REPLICA_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_1_1 | Oo_2_1 |
|---|
0016228
(v1.0) | Pias2 | NM_001164167,NM_001164168,NM_001164169,NM_001164170,NM_008602 | 17344
| 6.49135 | 5.5865 |
|
| NIA60mer |
| DBTMEE | Array_Feature_Number | Oligo_Parent_Clone | Unigene | Gene | Entrez | U_Cluster | NAP_Cluster | Gene_U/NAP | TIGR | SwissProt | EST | Oo | 1C | 2C | 4C | 8C | Morula | Blastocyst |
|---|
0016228
(v1.0) | 6850 | H3035G01-3 | Mm.6370 | Miz1 | 17344
| U018914 | NAP018092-001 | 6330408K17Rik;Miz1; | TC932587,TC848892,TC827274 | O54987 | U,F,2 | 2.5442 | 2.6934 | 2.4276 | 2.6707 | 2.4827 | 2.5601 | 2.6588 |
0016228
(v1.0) | 10336 | H3094D05-3 | Mm.6370 | Miz1 | 17344
| U018914 | NAP100923-001 | 6330408K17Rik;Miz1; | TC887276 | O54987 | U,F,2 | 2.4908 | 2.4418 | 2.0408 | 1.913 | 2.092 | 2.1204 | 1.9981 |
|
| MOE430 |
| DBTMEE | Probe_ID | Unigene | Gene | RefSeq | Entrez | Oo | 1C | 2C | 8C | Blastocyst |
|---|
0016228
(v1.0) | 1426456_a_at | Mm.6370 | Pias2 | NM_001164168.1,NM_008602.4,NM_001164167.1,NM_001164170.1,NM_001164169.1 | 17344
| 396.175 | 241.15 | 433.55 | 188.725 | 275.45 |
0016228
(v1.0) | 1419454_x_at | Mm.6370 | Pias2 | NM_001164168.1,NM_008602.4,NM_001164167.1,NM_001164170.1,NM_001164169.1 | 17344
| 54.225 | 76.05 | 46.75 | 0 | 152.2 |
0016228
(v1.0) | 1428725_at | Mm.6370 | Pias2 | NM_001164168.1,NM_008602.4,NM_001164167.1,NM_001164170.1,NM_001164169.1 | 17344
| 284.775 | 404.425 | 0 | 180.425 | 457.475 |
|
| Agilent44K |
| DBTMEE | Probe_ID | Gene | Acc | Entrez | Oo1_xFold | Oo1_pVal | Oo1_Flag | Oo2_xFold | Oo2_pVal | Oo2_Flag |
|---|
0016228
(v1.0) | A_51_P339858 | Miz1 | NM_008602 | 17344
| 1 | 1 | P | 0.861 | 0.391 | P |
0016228
(v1.0) | A_52_P470306 | Miz1 | NM_008602 | 17344
| 1 | 1 | P | 1.24 | 0.277 | P |
|
| Proteome |
N/A |
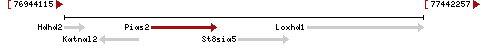
 12 Terms
12 Terms