| Gene | Apc | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms | CC1 Min | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | adenomatosis polyposis coli | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Locus | NM_007462 UCSC: chr18:34380638-34481844(+) OLP: chr18:34380638-34481844(+) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DB links | 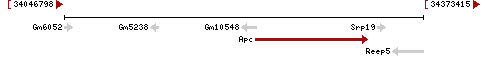 Entrez 11789 MGI 88039 Ensembl ENSMUSG00000005871 Vega OTTMUSG00000035932 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Feature | protein-coding | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO_BP |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO_MF |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| GO_CC |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ver2_FPKM | see DBTMEE:2002551 (v2.0) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SPR_Histone_Erkek_5KaroundTSS | N/A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SPR_Histone_Erkek_2KaroundTSS | N/A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SPR_Methy_Kobayashi_5KaroundTSS | N/A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SPR_Methy_Kobayashi_2KaroundTSS | N/A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_OocyteTo4C_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_Illumina_Oo_1C_Sperm_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_Partheno1C_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_Illumina_Oo_p1C_Sperm_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_Partheno4C_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Illumina_Oo_2C_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SOLiD_SC_Oo_REPLICA_FPKM_gene_guided |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NIA60mer |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MOE430 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Agilent44K |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Proteome | N/A |
 58 Terms
58 Terms