DBTMEE:0005704
| Gene | Dab2 |
| Synonyms | D15Wsu122e
p96 |
| Description | disabled 2, mitogen-responsive phosphoprotein |
| Locus | NM_001008702 UCSC: chr15:6249789-6390709(+) OLP: chr15:6249789-6390709(+)
NM_023118 UCSC: chr15:6336748-6390709(+) OLP: chr15:6336748-6390709(+)
NM_001102400 UCSC: chr15:6336748-6390709(+) OLP: chr15:6336748-6390709(+)
NM_001037905 UCSC: chr15:6336748-6390709(+) OLP: chr15:6336748-6390709(+)
|
| DB links |
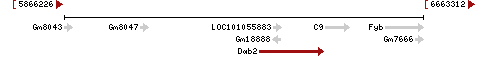
Entrez 13132
MGI 109175
Ensembl ENSMUSG00000022150
Vega OTTMUSG00000033430
|
| Feature | protein-coding |
| GO_BP |
-
 41 Terms
cell morphogenesis involved in differentiation (GO:0000904)
in utero embryonic development (GO:0001701)
positive regulation of receptor recycling (GO:0001921)
positive regulation of protein phosphorylation (GO:0001934)
transport (GO:0006810)
endocytosis (GO:0006897)
receptor-mediated endocytosis (GO:0006898)
pinocytosis (GO:0006907)
apoptotic process (GO:0006915)
activation of JUN kinase activity (GO:0007257)
multicellular organismal development (GO:0007275)
endoderm development (GO:0007492)
excretion (GO:0007588)
positive regulation of epithelial to mesenchymal transition (GO:0010718)
positive regulation of pathway-restricted SMAD protein phosphorylation (GO:0010862)
protein transport (GO:0015031)
Wnt receptor signaling pathway (GO:0016055)
cell differentiation (GO:0030154)
negative regulation of cell growth (GO:0030308)
positive regulation of cell migration (GO:0030335)
positive regulation of transforming growth factor beta receptor signaling pathway (GO:0030511)
negative regulation of protein binding (GO:0032091)
positive regulation of proteasomal ubiquitin-dependent protein catabolic process (GO:0032436)
positive regulation of transcription elongation from RNA polymerase II promoter (GO:0032968)
negative regulation of apoptotic process (GO:0043066)
positive regulation of cell adhesion (GO:0045785)
positive regulation of endocytosis (GO:0045807)
negative regulation of transcription, DNA-dependent (GO:0045892)
positive regulation of transcription, DNA-dependent (GO:0045893)
clathrin coat assembly (GO:0048268)
positive regulation of SMAD protein import into nucleus (GO:0060391)
negative regulation of androgen receptor signaling pathway (GO:0060766)
negative regulation of ERK1 and ERK2 cascade (GO:0070373)
cellular response to epidermal growth factor stimulus (GO:0071364)
cellular response to transforming growth factor beta stimulus (GO:0071560)
negative regulation of canonical Wnt receptor signaling pathway (GO:0090090)
positive regulation of substrate adhesion-dependent cell spreading (GO:1900026)
positive regulation of Wnt receptor signaling pathway, planar cell polarity pathway (GO:2000096)
positive regulation of clathrin-mediated endocytosis (GO:2000370)
positive regulation of early endosome to late endosome transport (GO:2000643)
positive regulation of integrin-mediated signaling pathway (GO:2001046)
|
| GO_MF |
-
 10 Terms
|
| GO_CC |
-
 8 Terms
|
| KEGG |
-
 1 Terms
|
| Ver2_FPKM |
see DBTMEE:2005629 (v2.0)
|
| SPR_Histone_Erkek_5KaroundTSS |
N/A |
| SPR_Histone_Erkek_2KaroundTSS |
N/A |
| SPR_Methy_Kobayashi_5KaroundTSS |
N/A |
| SPR_Methy_Kobayashi_2KaroundTSS |
N/A |
| SOLiD_OocyteTo4C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | SPR_FPKM | Oo_FPKM | 1C_FPKM | 2C_FPKM | 4C_FPKM | MEF_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 31.834 | 41.0353 | 6.92405 | 7.50779 | 12.0625 | |
|
| SOLiD_Illumina_Oo_1C_Sperm_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | SPR_FPKM | 1C_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 33.2853 | 14.109 | 44.7704 |
|
| SOLiD_Partheno1C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | 1C_FPKM | p1C_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 32.4898 | 40.6233 | 39.1141 |
|
| SOLiD_Illumina_Oo_p1C_Sperm_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | SPR_FPKM | p1C_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 33.3047 | 13.9157 | 42.8991 |
|
| SOLiD_Partheno4C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | 2C_FPKM | 4C_FPKM | p4C_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 6.41059 | 7.24907 | 6.57632 |
|
| Illumina_Oo_2C_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_FPKM | 2C_FPKM |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 39.6504 | 26.45 |
|
| SOLiD_SC_Oo_REPLICA_FPKM_gene_guided |
| DBTMEE | Gene | RefSeq | Entrez | Oo_1_1 | Oo_2_1 |
|---|
0005704
(v1.0) | Dab2 | NM_001008702,NM_001037905,NM_001102400,NM_023118 | 13132
| 23.411 | 26.3289 |
|
| NIA60mer |
| DBTMEE | Array_Feature_Number | Oligo_Parent_Clone | Unigene | Gene | Entrez | U_Cluster | NAP_Cluster | Gene_U/NAP | TIGR | SwissProt | EST | Oo | 1C | 2C | 4C | 8C | Morula | Blastocyst |
|---|
0005704
(v1.0) | 4770 | G0104C04-3 | Mm.34248 | Dab2 | 13132
| U043423 | NAP009855-001 | Dab2; | TC936378,TC935218,TC935216,TC935215 | P98078 | U,F,2 | 3.1986 | 2.9858 | 2.7404 | 2.4707 | 2.8698 | 3.1627 | 3.0768 |
|
| MOE430 |
| DBTMEE | Probe_ID | Unigene | Gene | RefSeq | Entrez | Oo | 1C | 2C | 8C | Blastocyst |
|---|
0005704
(v1.0) | 1420498_a_at | Mm.240830 | Dab2 | NM_023118.5,NM_001037905.3,NM_001102400.1,NM_001008702.2 | 13132
| 548.85 | 660.725 | 312.575 | 542.1 | 1210.3 |
0005704
(v1.0) | 1423805_at | Mm.240830 | Dab2 | NM_023118.5,NM_001037905.3,NM_001102400.1,NM_001008702.2 | 13132
| 98.05 | 123.35 | 0 | 0 | 136.025 |
0005704
(v1.0) | 1430604_a_at | Mm.240830 | Dab2 | NM_023118.5,NM_001037905.3,NM_001102400.1,NM_001008702.2 | 13132
| 60.925 | 69.95 | 45.3 | 80.25 | 121.375 |
|
| Agilent44K |
| DBTMEE | Probe_ID | Gene | Acc | Entrez | Oo1_xFold | Oo1_pVal | Oo1_Flag | Oo2_xFold | Oo2_pVal | Oo2_Flag |
|---|
0005704
(v1.0) | A_51_P157994 | Dab2 | NM_023118 | 13132
| 1 | 1 | P | 1.21 | 0.307 | P |
|
| Proteome |
| DBTMEE | Gene | Entrez | GV_seqCount | MII_seqCount | Zygote_seqCount | GV_specCount | MII_specCount | Zygote_specCount |
|---|
0005704
(v1.0) | Dab2 | 13132
| 3 | 2 | 3 | 4 | 2 | 5 |
|
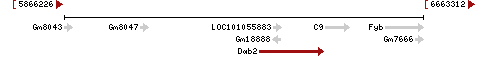
 41 Terms
41 Terms